Thomas Rattei’s work covers a wide spectrum of topics from bioinformatics, genome and metagenome analysis and systems biology. He has long-standing expertise in developing and applying computational methods for the interpretation of large-scale sequence information. The international reputation of his research group triggered their involvement in numerous international (meta-) genome sequencing and analysis consortia.
Thomas’ research activities not only cover individual, project-specific questions but also general problems in bioinformatics, computational infrastructure, and large-scale biological databases. Furthermore, his group develops novel, genome-based computational approaches for studying molecular inter-species interactions, such as between hosts and pathogens, between symbionts, or in microbial ecosystems.
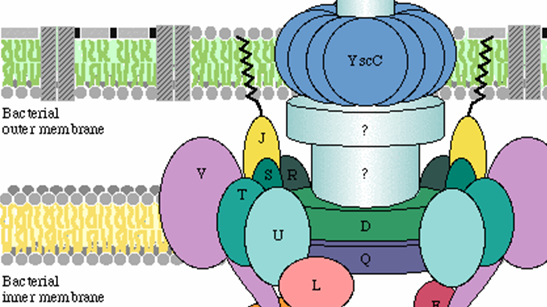
Thomas and his team maintain and develop internationally relevant resources in computational biology, such as the web portals phendb.org, vogdb.org and effectivedb.org for microbial trait prediction, virus orthologous groups and protein families, and bacterial secreted proteins and secretion systems.