Genotype-phenotype heterogeneity of bacterial colonies

Understanding the genotype to phenotype connection is one of the key open questions in biology.
Bioinformatics is an interdisciplinary field at the interface between computer science and the life sciences, with important applications in molecular medicine, healthy living, agriculture, biotechnology, green energy, and other areas. Not only does bioinformatics provide the information infrastructure on which these fields analyse their vastly growing repertoire of molecular data, but it is also becoming the data science and integration science that yield important new discoveries and thereby leverages society’s investment into the life sciences.

CUBE, the Division of Computational Systems Biology, is a group of bioinformaticians and computational biologists. We are interested in understanding of biological systems, ranging from single species to multi-species systems and ecosystems, based on data from large-scale bioanalytical methods. We develop, improve and apply computational methods for the interpretation of molecular information in biology, and establishes and analyses quantitative mathematical models.
CUBE is therefore highly engaged in basic research, in the translation of basic knowledge to medical and biological applications, and in the training of students on all educational levels. We support an innovative and open-minded, creative and collaborative atmosphere inside and outside our team, facilitate the exchange with the society and contribute to a vital and responsible research community in computational biology.


Understanding the genotype to phenotype connection is one of the key open questions in biology.
Many biological and evolutionary questions are related to the structure of the protein sequence space and its partitioning into substructures…
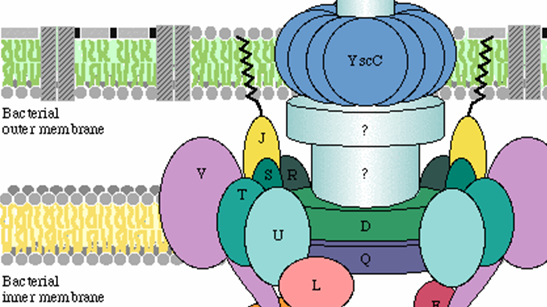
Annoying for most of us, but dangerous or even life threatening for particular persons: infections by pathogenic bacteria are never welcome.
We develop and provide access to various software and databases, including: PhenDB, VOGDB, NVT, HoloVir, SIMAP, EffectiveDB, PICA, Gepard, and GenSkew.

We operate and administer Life Science Compute Cluster (LiSC), a specialized high-performance computing infrastructure which is available to life scientists across the University of Vienna.
